The new product supports more breeds and returns higher accuracy through an updated reference panel. We have also made it easier to use for industrial scale applications.
We are excited to announce an update to our cattle imputation reference panel, which now includes over 50 breeds and 1800 animals. This better supports the demand we’ve had for all the most important breeds being utilized by the cattle industry today. Since launching our original panel, we’ve been busy optimizing our pipeline and adding new features to make low-pass sequencing more powerful and easier to use. We believe this platform is now the best high-throughput solution for building genomic breeding programs.
Our new panel now returns over 50 million variants that you can genotype with over 99% accuracy. This includes over 98+% of the sites for the existing arrays in production today, from the Illumina BovineHD to the SNP50. We support “Final Report” backwards compatibility for any available array manifest so that the data you generate can be seamlessly integrated into existing workflows. It also now characterizes indels as well as SNPs and multiallelic sites. We have exact matches at 98% of the SNPs in the ICAR200 parentage panel.
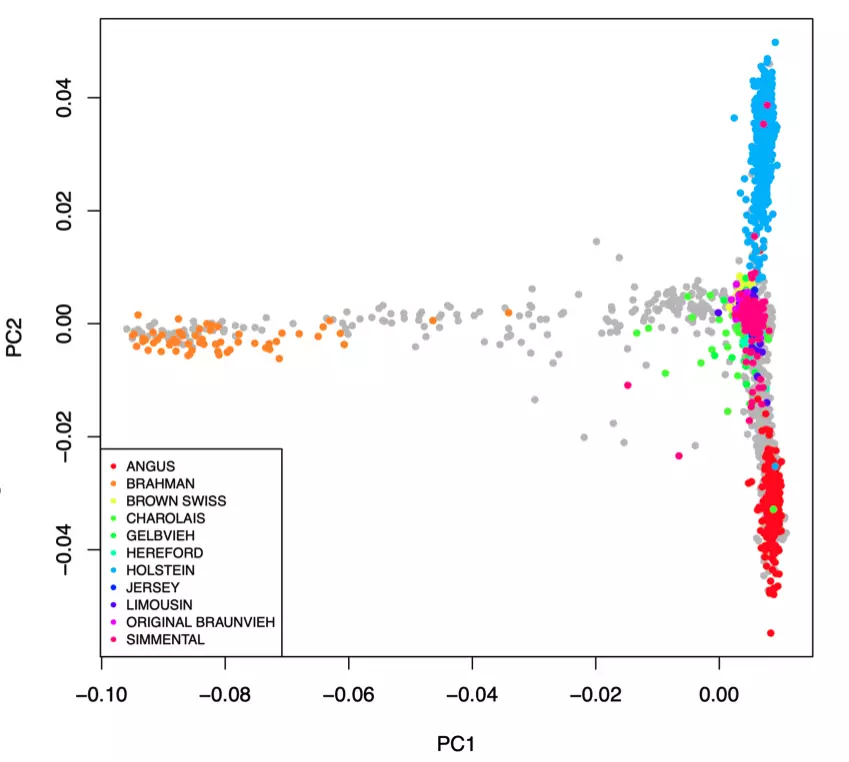
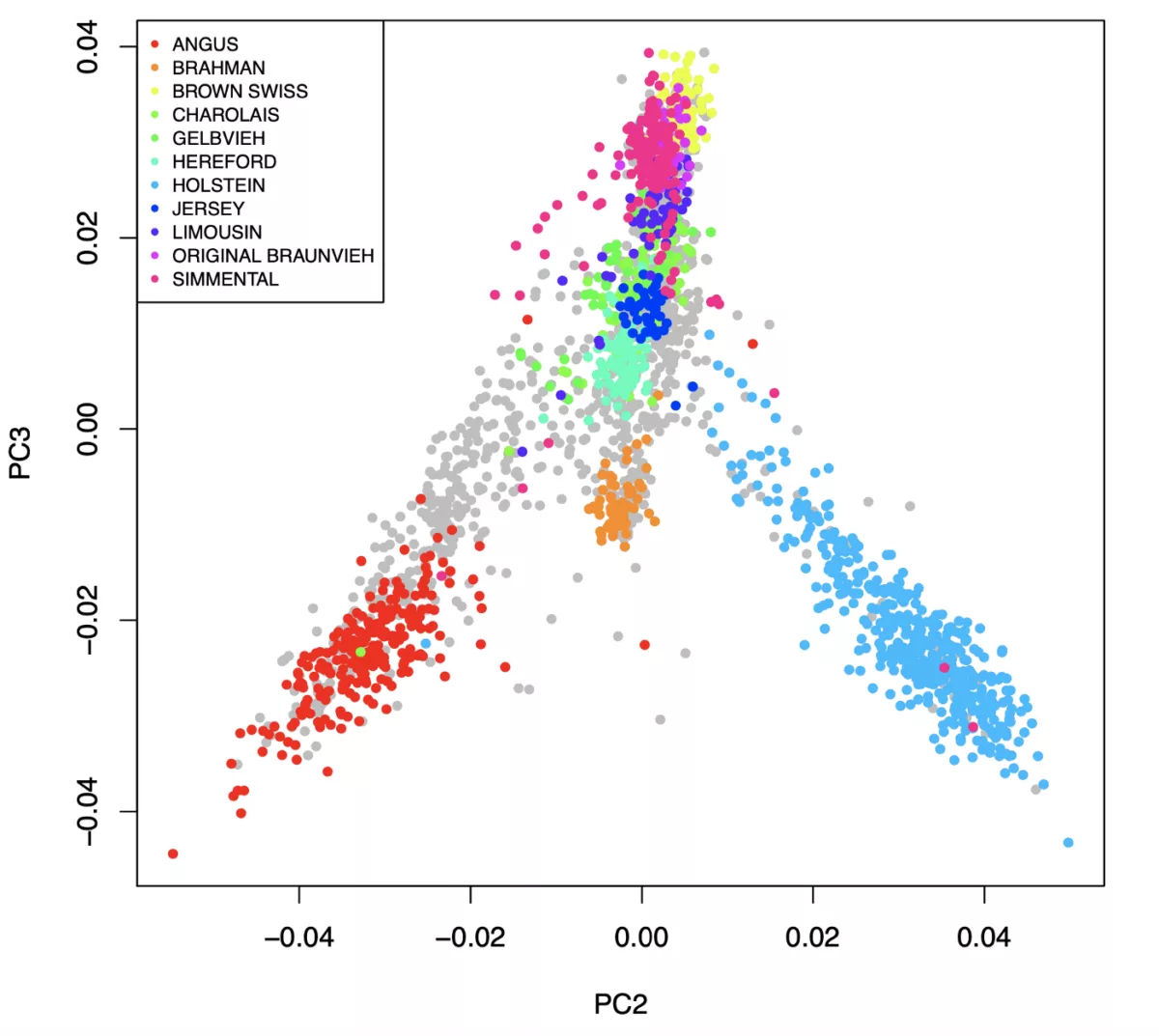
The key to successful low-pass genotyping is making sure the common haplotypes from a population are included in the reference. Our panel now has 9 breeds with over 40 samples each. Due to the large amount of sharing of common genetics amongst different European cattle breeds, this combined breed panel improves performance across the whole dataset (see below). Huge thanks to all the public research partners who released sequence data.