tl;dr: just use it as any other tool and see if it makes things work better. Unexpected insights may emerge along the way.
Blog
Tomaz Berisa - Jun 18, 2024
A practical approach to “AI” in genomics
Artificial Intelligence is a rapidly expanding field with applications across diverse industries, promising to transform how we live and work. The cynic would say “That’s great, but atm it’s a cool digital assistant and image generator”. Reconciling these opposing instincts such that one appropriately calibrates for the strategic importance of these technologies — while maintaining a healthy grounding in reality — is not an easy task.
At Gencove, we’ve observed non-trivial improvements in performance and productivity from using publicly available and in-house deep neural networks, so we wanted to share how we approach the topic of setting strategy and building, while separating utility from hype. We also provide several examples of projects we found insightful as we navigate the space.
The process
“When in doubt, apply the product process.”
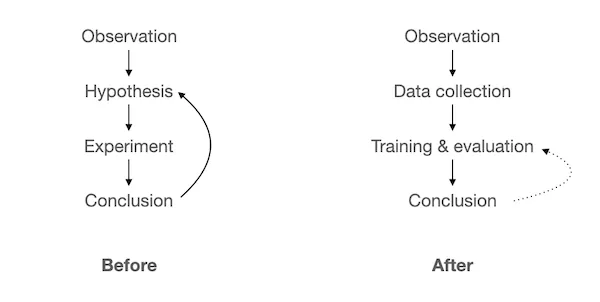
Historically, many product tickets at Gencove were mini PhD projects taking longer than we’d like, but rigorous application of our product process (heavily influenced by The 4 Big Risks and Shape Up) clearly identified situations where applying deep neural networks was the most efficient approach and not chasing a shiny object — “AI”.
This process leads us to something much more valuable than just well-performing solutions to the specific problems at hand: a more general problem solving approach where we provide examples and let the model figure it out, instead of painstakingly iterate through hypotheses until we find something that works.
Kudos to our product team for setting up an effective and simple process that consistently keeps our eye on the ball — we consider this one of our key advantages as a company.
Emergent properties: strategy
“The best outcome of a product ticket is to influence broader strategy”
We also noticed that some of the solutions generalize very nicely outside of the initial problem’s scope — ultimately rising to the level of a strategic theme at Gencove.
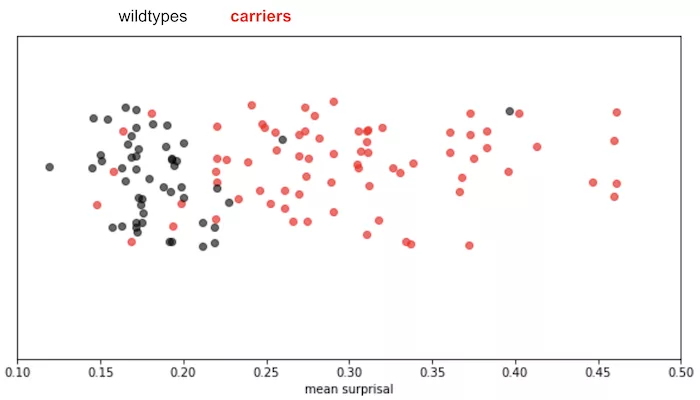
For example, we were tasked with using short-read sequencing to call a rare ~800 bp insertion that appears in carriers of a specific disease, where traditional methods were having issues calling the variant. We went ahead and trained a small GPT (a type of LLM) on sequencing reads from wildtype samples and assessed the model’s surprise when attempting to make predictions on reads coming from wildtype and carrier samples in the test set. As shown in the figure below, we can see that the model is clearly surprised by the carrier samples more than it is surprised by wildtypes:
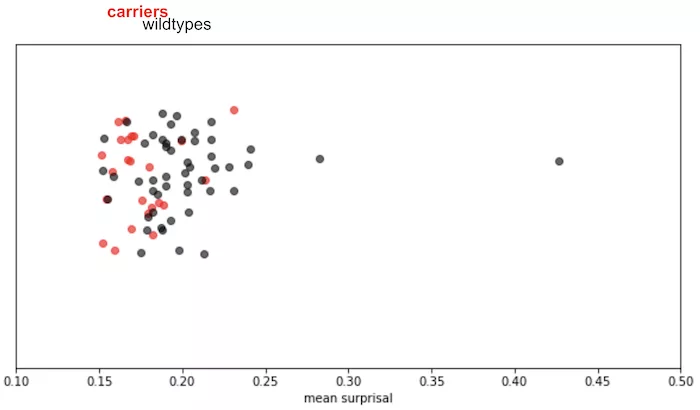
Furthermore, we observed that fine-tuning on reads from a subset of carrier samples results in a model that is no longer surprised by remaining carriers in the test set:
This shows that the model was indeed detecting systematic differences between reads from the carries and wild-type individuals.
The presented model is a fun proof-of-concept, but extrapolating the result to a broader model leads us to what we like to call a generalized hypothesis-free outlier detector, i.e., a model that can be:
- Trained on samples representing what is considered “normal” (thus far)
- Used to detect unexpected samples, given what was considered normal
- Fine-tuned (or not) on the unexpected samples to mark them as “normal” moving forward
The most valuable aspect of the method is that it doesn’t need to be told what to look for or how to identify outliers, rather just to identify things that do not look like what it has already seen.
My favorite (albeit quite aspirational) analogy here is Tesla Autopilot. It is a machine learning system that is constantly collecting vast amounts of observations from the real world and updating its ability to detect the edge cases and operate in such an environment. In a sense, Tesla is sampling the variation of traffic information and control mechanisms in the wild (which is surprisingly complex), while we are sampling genetic variation across populations and species.
We see at least three applications of such a model continuously running in the background of industrial-scale sequencing analytics operations (our bread and butter at Gencove):
- “Looking for hidden value” — identify the outliers when looking at clinical trial data in humans or breeding program for plants and animals
- “Saving time” in terms of assisting data analysis and/or cleanup
- Quality control at scale
We codenamed this approach Watchdog and are quite excited about its applications in the field.
This section outlines two more examples of solutions we found insightful.
Large insertion caller
The GPT approach to resolving that 800 basepair insertion (from the previous section) was a fun project that yielded far more insight than we bargained for, but we were confident we could do better at the specific task of calling that specific variant. Taking inspiration from DeepVariant, we trained a CNN with thousands of IGV plots of the locus appropriately labeled as wildtype or carrier, and the model was over 99% accurate. All this was an unoptimized afternoon of work, as opposed to a much longer iterative hypothesis-driven process.
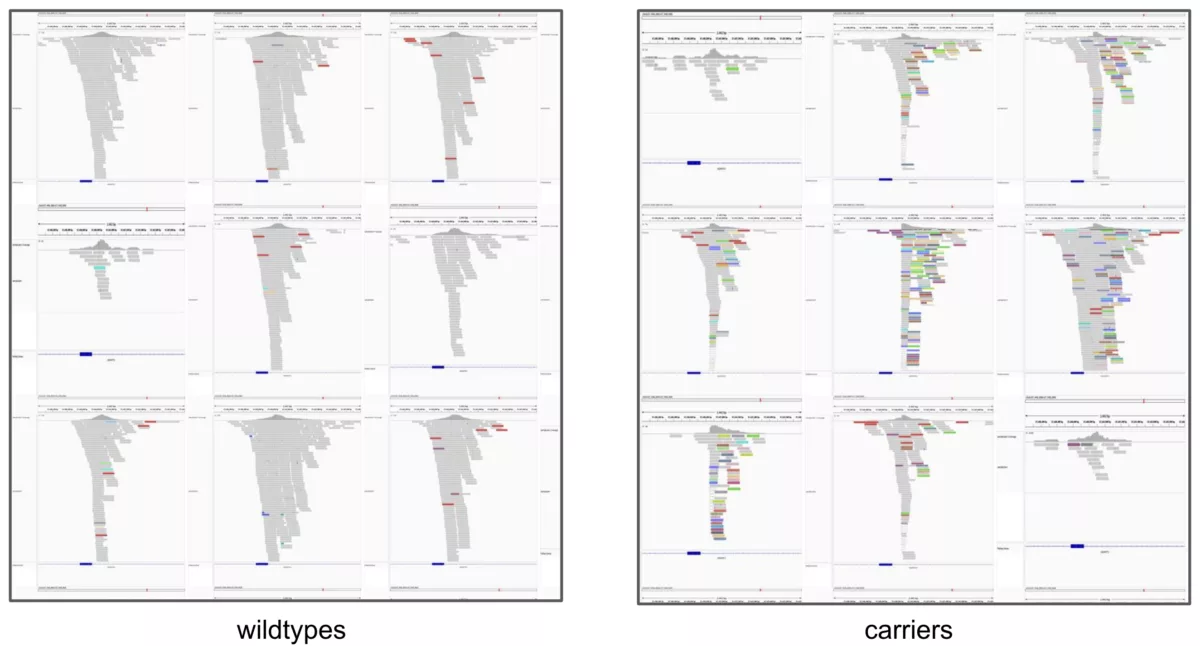
As a sanity check, visual inspection of the IGV plots clearly demonstrates the two classes “look different”, i.e., why it’s not particularly surprising that a CNN can differentiate between them:
Sample mixture analysis
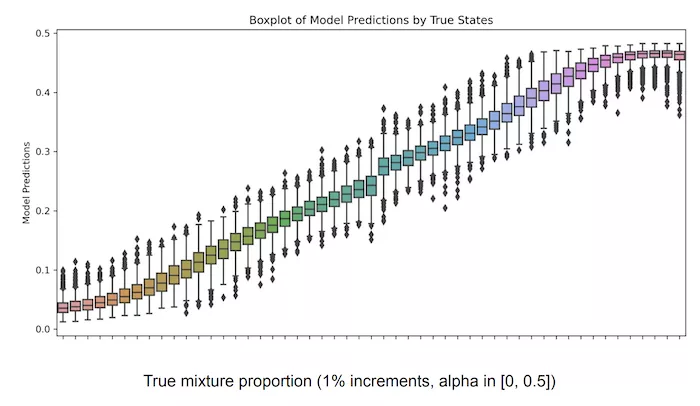
A common problem across many applications of genomics is detecting sample mixtures and quantifying the respective mixture proportions. Examples that immediately come to mind are liquid biopsies, NIPT, and forensics. This problem is hard enough with high-coverage sequencing data, but becomes significantly more difficult with low-coverage data from individuals of mixed ancestries. To bypass the pain, we decided to train a relatively small MLP with sequencing reads from a modest number of two-sample mixtures with known mixture fractions and see if it can predict the correct mixture proportions. At this point it’s not a surprise that the model performed quite well:
We hope this has been an informative read and are excited to continue working on these and other interesting classes of problems in genomics — stay tuned for more from the team. We would also be delighted to keep the discussion going and look forward to hearing thoughts about these and other applications from the community.
If you’re interested in building these types of systems together — just reach out to me directly at tomaz@gencove.com