Gencove’s low-pass whole genome sequencing and analysis platform is a hardware-agnostic high-throughput and cost-effective software layer. Gencove significantly reduces the cost per sample in three ways. First, by miniaturizing DNA requirements and library preparation. Second, by allowing customers to sequence at lower coverages and then applying the Gencove’s software to impute against species-specific haplotype reference genomes. Lastly, by delivering high-quality variant calling, the Gencove platform enables ongoing downstream analysis.
To offer another option for scientists and companies seeking whole genome sequencing and analysis, Gencove evaluated the ability to process sequencing data derived from the AVITI™ System in two studies.
Validation study design
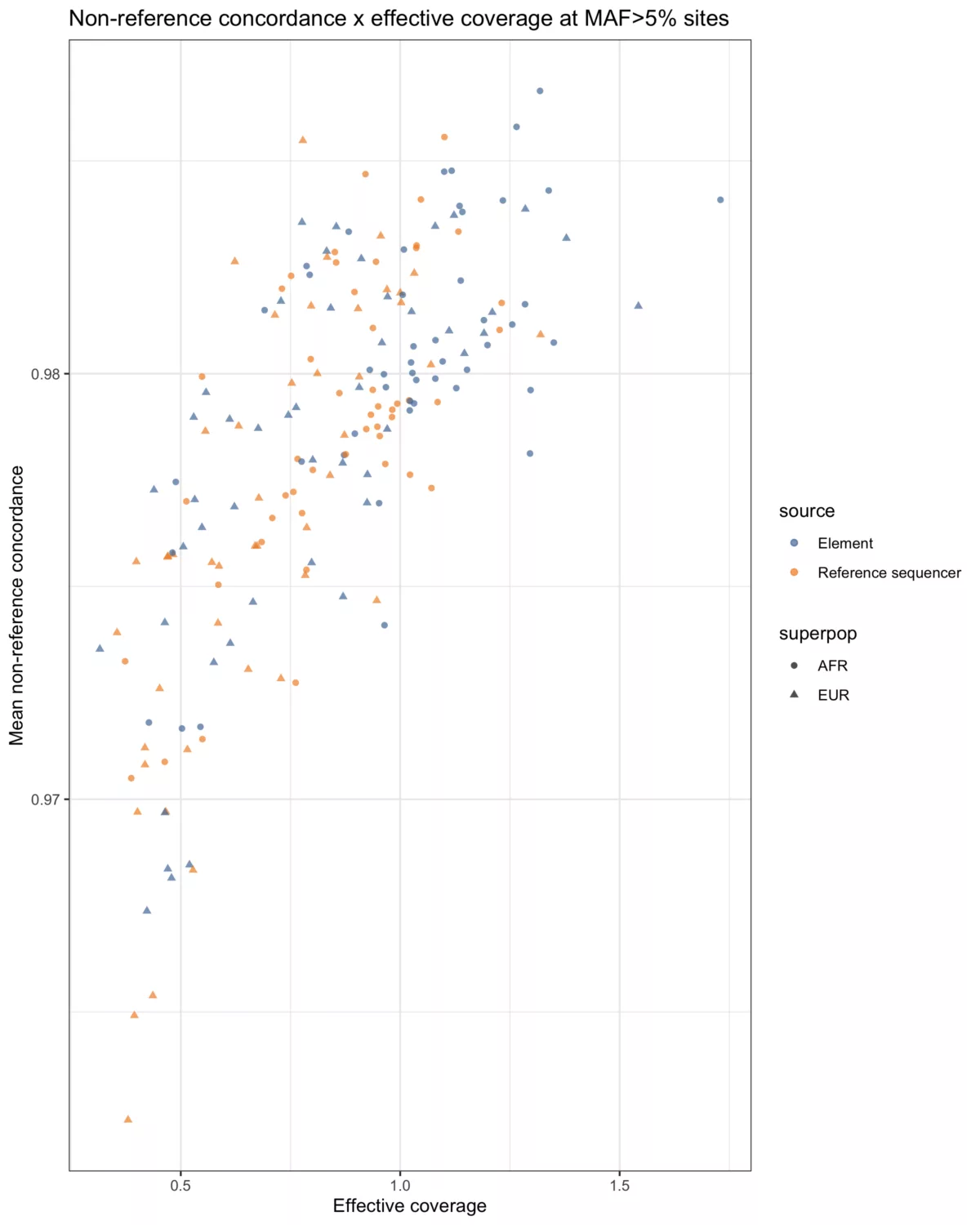
The validation design mimics the Li et al. study (2021) published in Genome Research where libraries were prepared from 48 European and African samples from the 1000 Genomes Project using KAPA library prep kits. Samples were pooled and sequenced on a reference sequencer and the AVITI SystemThen the data were imputed in a leave-one-out manner to the 1000 Genomes Phase 3 reference panel.
Validation results
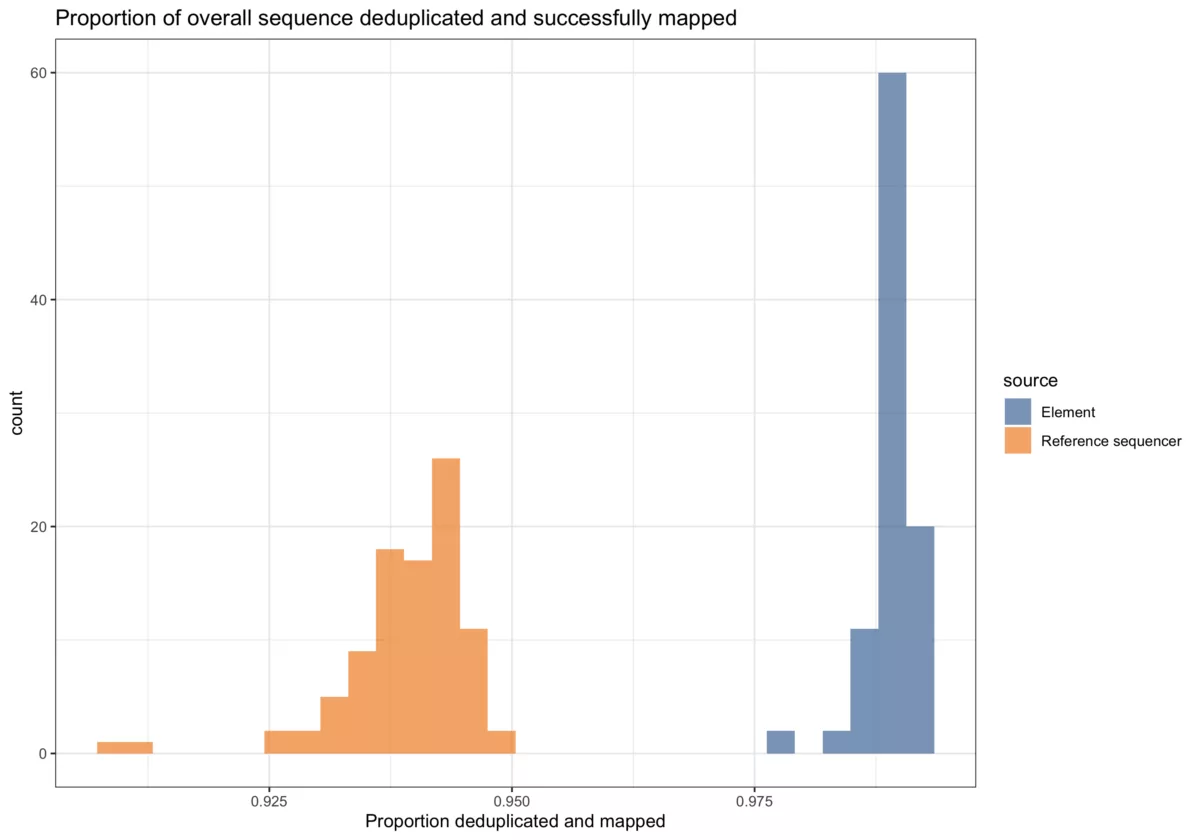
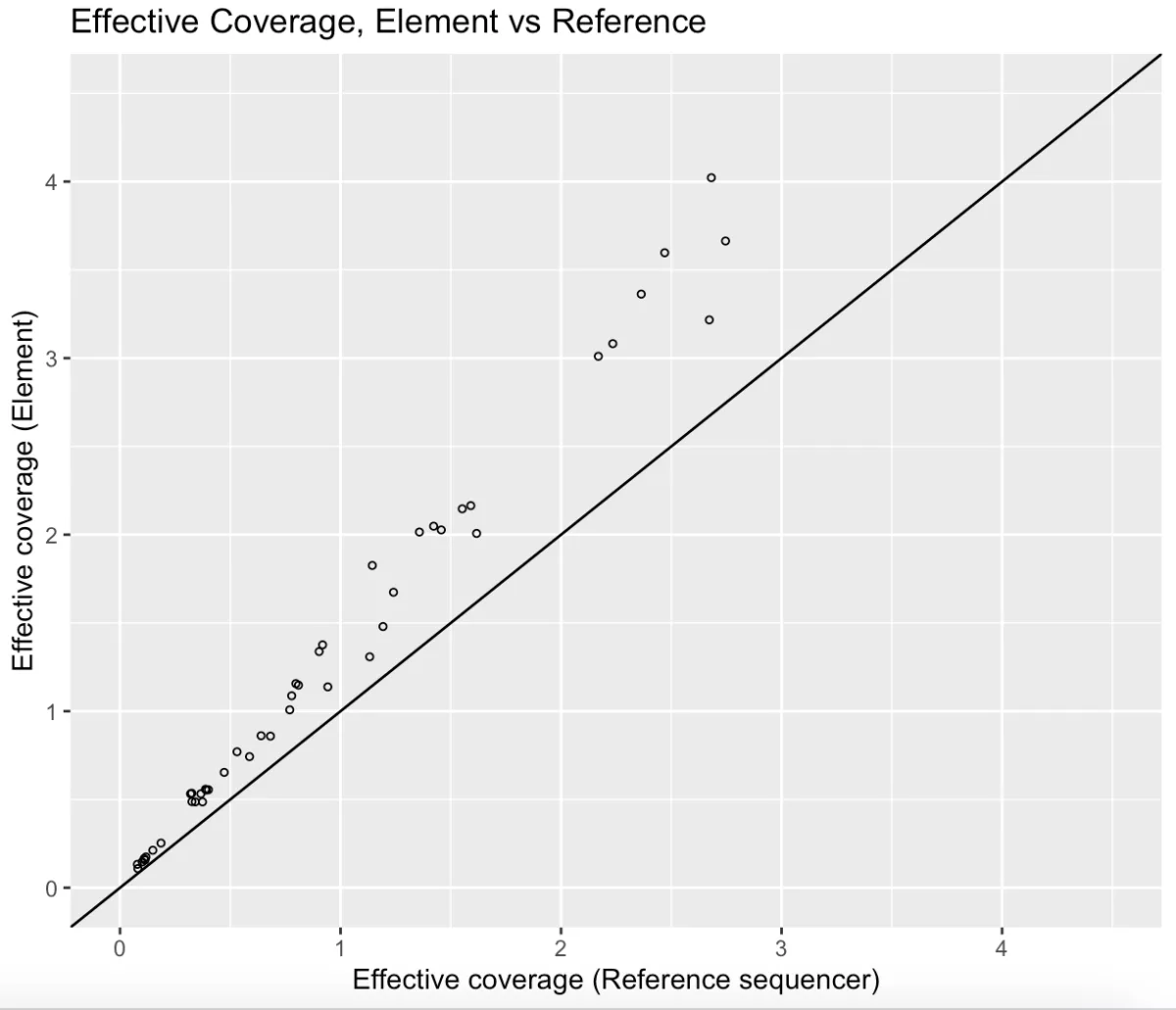
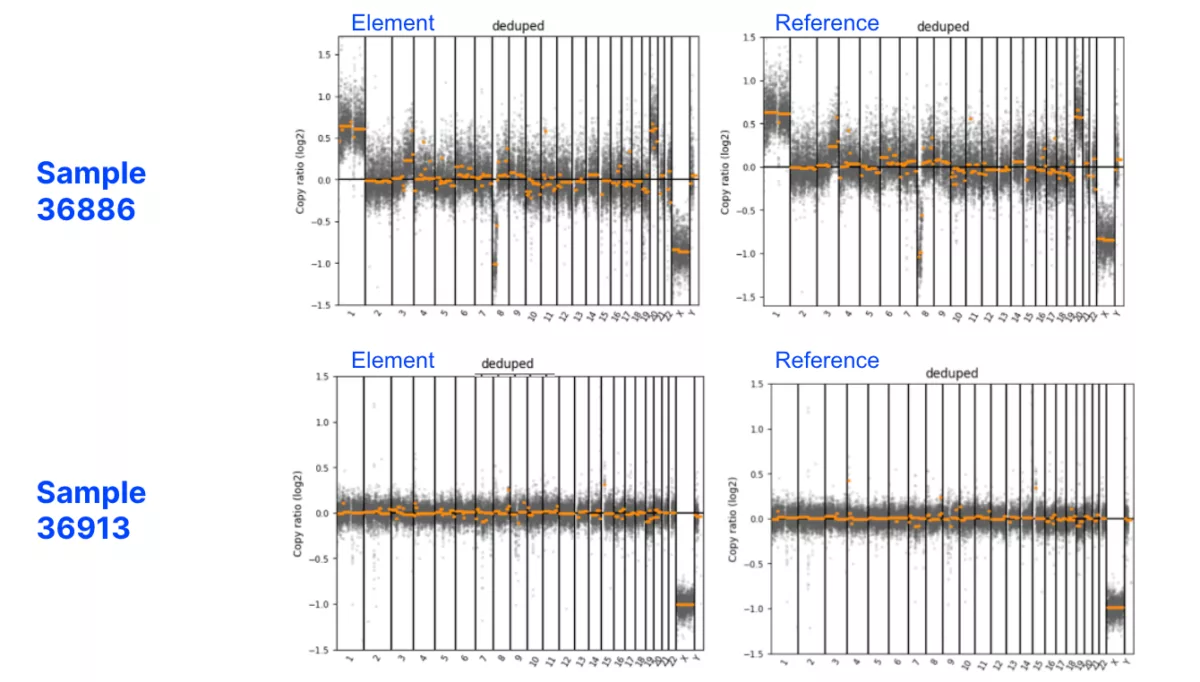
The AVITI System is concordant with the reference sequencer for low-pass whole genome sequencing.