New Gencove Feature Released
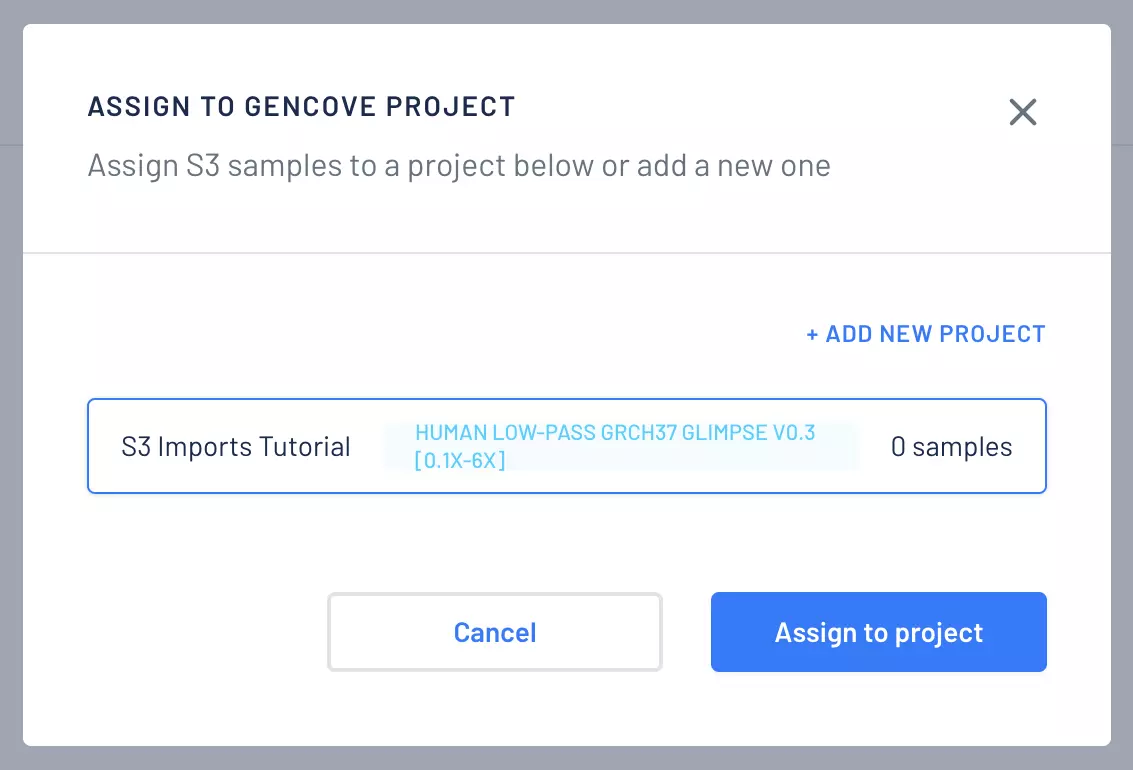
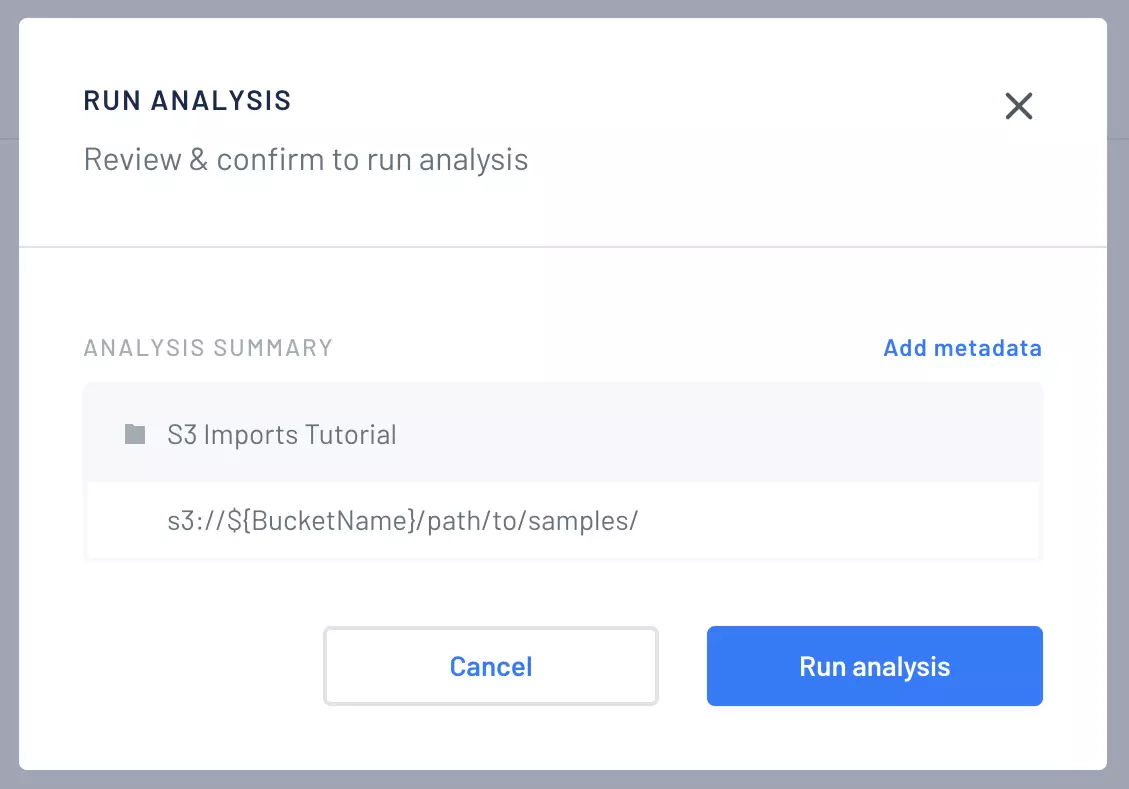
At Gencove, we are committed to making genome sequencing more accessible. To that end, we are pleased to announce the ability to use Amazon S3 directly as an import origin in addition to supporting uploads through our CLI, our API, or from BaseSpace. The new feature saves time because samples can now be imported directly from an S3 bucket without the need to download and then upload again. In this post, we show how to import samples from a S3 URI to a project.
How It Works
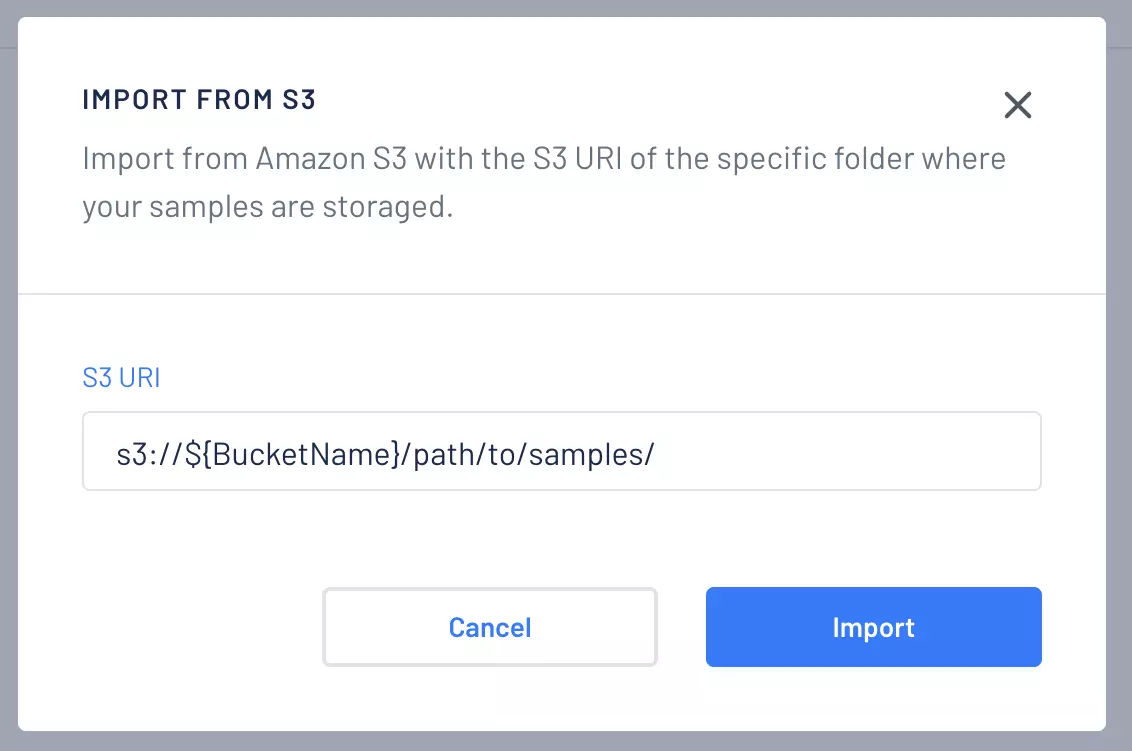
To be able to import samples first you need to have a working S3 connection, then you can provide the S3 URI of the folder (i.e., prefix) where your samples are located.