In order to leverage the information that improves crop productivity and profitability, breeders need cost-effective solutions that will provide them with genome wide, highly accurate information. Existing targeted genotyping by sequencing (tGBS) methods are only able to provide insight into limited numbers of targets and have challenges with call rate, designability and accuracy. Gencove has previously developed solutions for livestock that address this, and now we’re excited to demonstrate the application of these technologies in plants.
Blog
Jesse Hoff, Agrigenomics Product Manager - Jul 10, 2023
Exploring the power of low-pass WGS with target capture in plant breeding
A new solution for genetic insights
Low-pass whole genome sequencing (lp-WGS) with target capture and imputation offers a solution to provide genome wide information and highly accurate calls at the most important loci in the genome. In addition, this new approach has the advantage of being affordable and high throughput.
Gencove and Twist Biosciences partnered to pressure test the utility of lp-WGS plus target capture and imputation for a major crop species, soybeans, demonstrating that lp-WGS with target capture works well in plants and covers a large panel. Here we generated high coverage data on a ~70kb panel. Target genes were chosen based on their previous focus in academic efforts on plant gene editing.
What is low-pass whole genome sequencing with target capture?
Low-pass whole genome sequencing with target capture and imputation combines a number of different, highly effective sequencing approaches in a single assay. With lp-WGS, the genome is shotgun sequenced at a low coverage (typically <1x), delivering genome-wide data which is then imputed to a haplotype reference panel to provide genome-wide genotype calls. In addition, target capture techniques are employed to capture specific regions of interest by the design and use of custom probes to target desired genomic regions. These probes selectively bind to and capture the regions of interest from the whole-genome sequencing library, allowing for a higher-fidelity analysis at key high-value loci.
The combination of low-pass whole-genome sequencing and target capture allows us to efficiently analyze and study specific genomic regions while still obtaining a broad overview of the entire genome. This approach is particularly useful when investigating specific genomic regions associated with particular traits, as it provides a cost-effective and targeted solution for genomic analysis while still supporting breeding objectives.
Project protocol
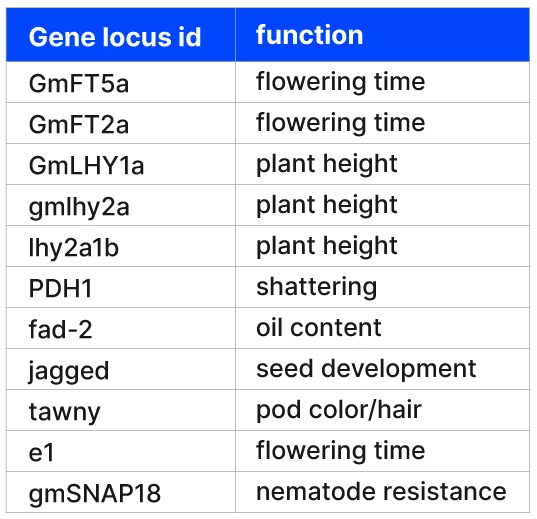
Alongside providing soybean DNA samples, Gencove identified 11 soybean gene targets of interest. These genes have previously been the subject of gene editing experiments. Therefore projects involving discovery of standing natural variation, as well as ongoing management of gene editing events could be of interest to a breeding program. The team at Twist sequenced the samples to 1x coverage with lp-WGS, and at 30x at the 11 target sites. Gencove then performed variant calling, low-pass imputation and QC on output data, and returned results on the success of capture at the 11 target sites.
Results
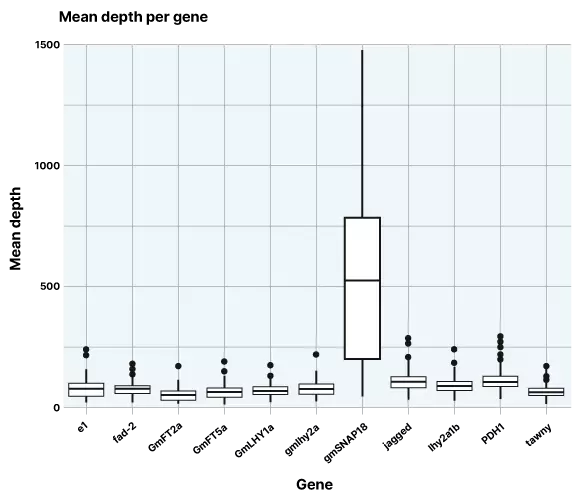
Figure 1. Mean depth across 11 target genes
Data produced from the project (figure 1) show consistently high depths for the target genes, with a median depth across target genes of 74.66, which demonstrates the reliability of the lp-WGS with target capture approach. As is often the case in plant genomics, we observed artifactual coverage in gmSNAP18 that stands out in the above chart, this is due to it likely being a complex locus and so caution is needed in interpretation. However, this result did not impact the overall performance of the rest of the panel or impact further downstream analysis.
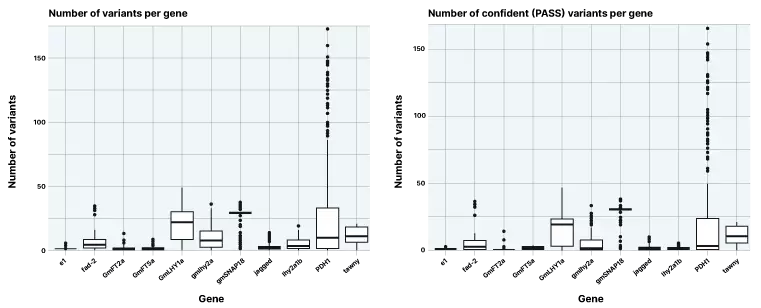
Figure 2. Number of novel variants identified across 11 target genes
The project focused variant discovery on the specific genes of interest. Critically, across the 11 target genes we saw robust identification of variants, with the number of novel high confidence SNPs and indels that were discovered for each gene compared to the reference panel shown in the figure above.
A lp-WGS sequencing approach with target capture allows us to identify both SNPs and indels. Our assay and results reflect the range of standing variation observed in different genes in soybean. Additionally, the modeling for SNP discovery with high coverage short reads allows for informative filtering.
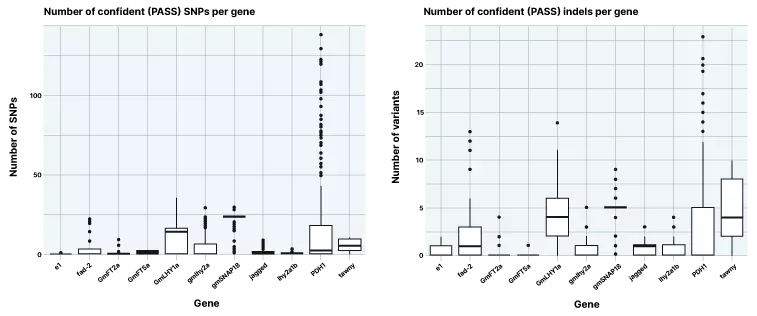
Figure 3. Number of confident SNPs and indels identified across 11 target genes
Short reads with high coverage are suitable for discovery and characterization of SNPs and indels. While there are fewer indels discovered on average, these represent an abundant quantity of standing variation in commercial soy populations.
Excellent and repeatable coverage and enrichment
The combination of lp-WGS and target capture is an exciting and effective offering for plant breeders. In a single, reliable molecular reaction, lp-WGS plus target capture enables a cost-effective solution for direct observation of the known and unknown genetic variation across the whole genome. The panel was easy to design and large, powered by the quality and speed of Twist’s platform. Set up took a few weeks, and further iterations of the assay typically do not require protocol optimization. This enables new targets to be added as your breeding program evolves. Meanwhile, the low pass data provides the reliable backbone for genome wide genomic evaluation.
Additionally this approach can be run on any sequencing platform; you can check out our recent webinar with Element Biosciences to learn more about our projects on the AVITI System.
Gencove is continually innovating to bring genomic information to the forefront of advancing global health and sustainability. Understanding the genetic traits of crop species empowers breeders to make informed decisions in crop management, and variety selection for profitability and productivity
Gencove would like to thank the University of Minnesota for their collaboration and sharing these samples.