Molecular breeding has made significant strides over the past decade and is driving key discoveries across many species. Funding for genomics and genetics-powered breeding programs in both plants and animals by the USDA alone is well into the tens of millions of dollars per year.
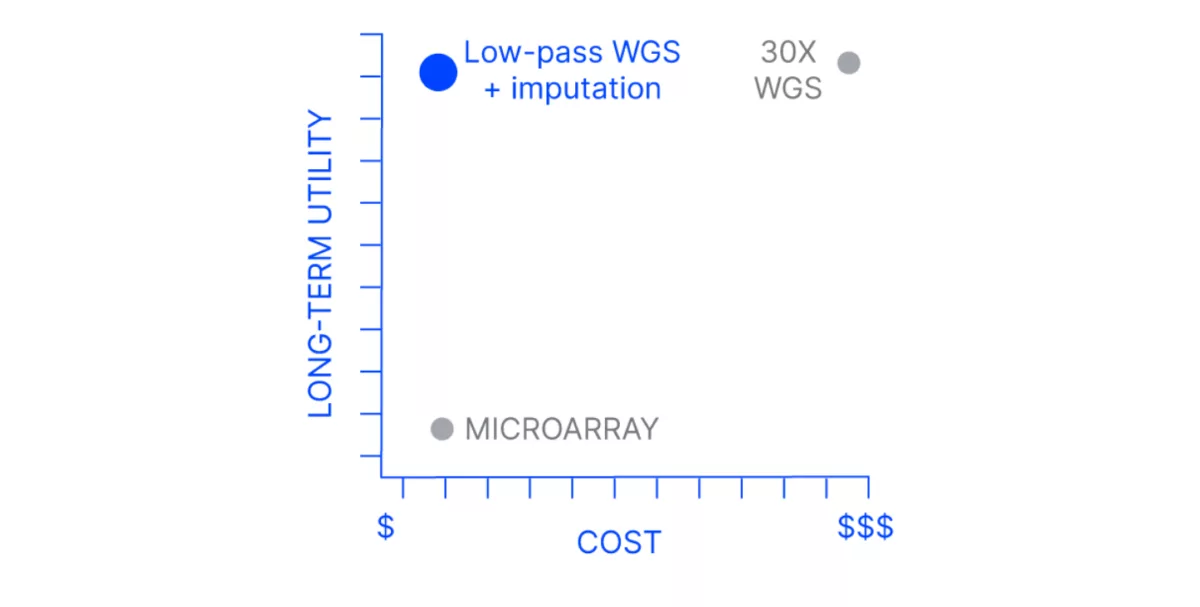
Drivers of the investment are the increased utility and the decreasing cost of sequencing technologies. Sequencing is the foundation of major milestones such as completing a complex genome from telomere to telomere and identifying genetic variation in dispensable genes of a pangenome. However, the cost of the gold standard 30X whole genome sequencing (WGS) coverage for variant detection projects prohibits its use as a routine, high-throughput genotyping tool.
On the other end of the spectrum, microarrays are inexpensive on a cost-per-sample basis after the initial time and financial investment is made to develop the chip. In breeding programs where known markers are used to power genomic selection and no updating or development of new markers is expected, a high-density microarray may provide enough information for breeding decisions.
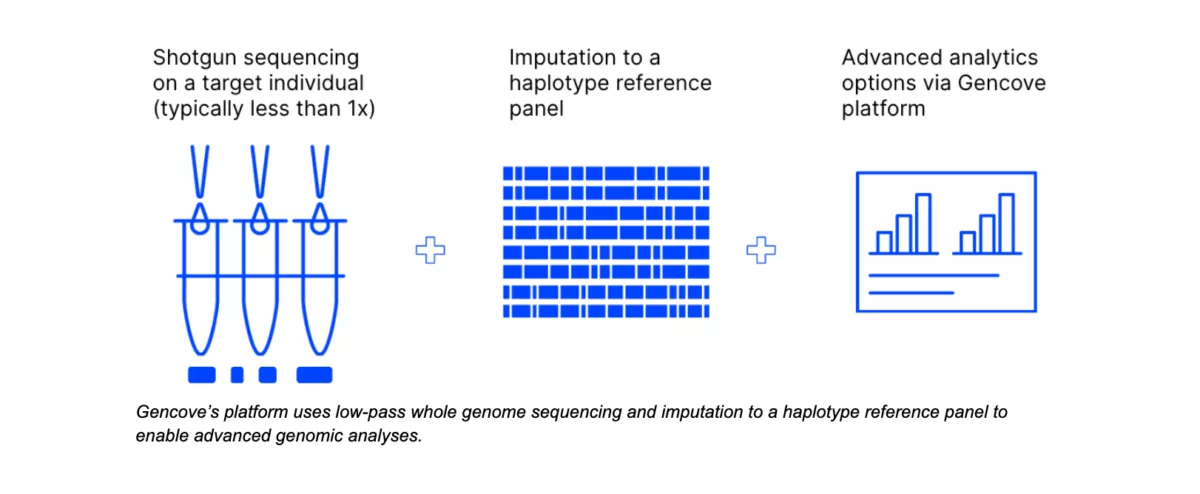
However, if you’d prefer a pipeline that genotypes your known key markers and also provides genome-wide information for millions of other variants for future exploration, Gencove’s low-pass whole genome sequencing and analysis platform is a powerful, cost-effective, high-throughput, future-proofed solution.