Improving genetics for traits such as higher yield, drought tolerance, and disease susceptibility depends on translating genomic information into screening and prediction tools for fast, efficient, and sustained breeding progress. From using pedigree information to identifying key structural variants via sequencing, the tools used to make informed decisions based on genetic variation have advanced tremendously over the last decade. However, while technology has made large strides, continued use in large breeding programs requires genomic information to be useful, fast, cost-effective, and easy to implement.
Blog
Jesse Hoff, Agrigenomics Product Manager - Jun 15, 2022
A Modern Approach to Genotyping for Plant and Animal Breeding
Sequencing: the powerhouse of genomic studies
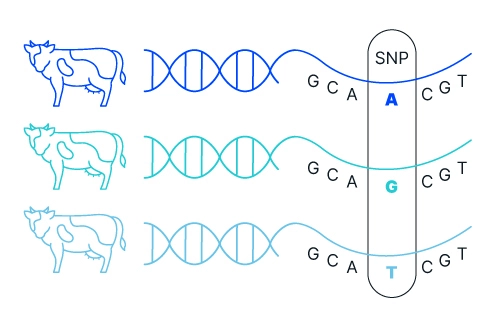
DNA sequencing, especially of the whole genome, has emerged as one of the most useful tools for understanding genomes and genetic variation amongst and between species. Whole genome sequencing is used for generating high-quality reference genomes to serve as a benchmark for comparisons to identify genetic variants from SNPs, to indels, to large structural variants. Sequencing is used for QTL mapping efforts to tie genetic variation to specific traits targeted by breeding programs. Sequencing has also been used to identify and develop the markers to be used for genotyping tools like microarray.
There’s no question that sequencing is a powerhouse of utility for genetics in breeding. However, does it meet the requirements to become a routine tool for genomic selection in breeding programs?
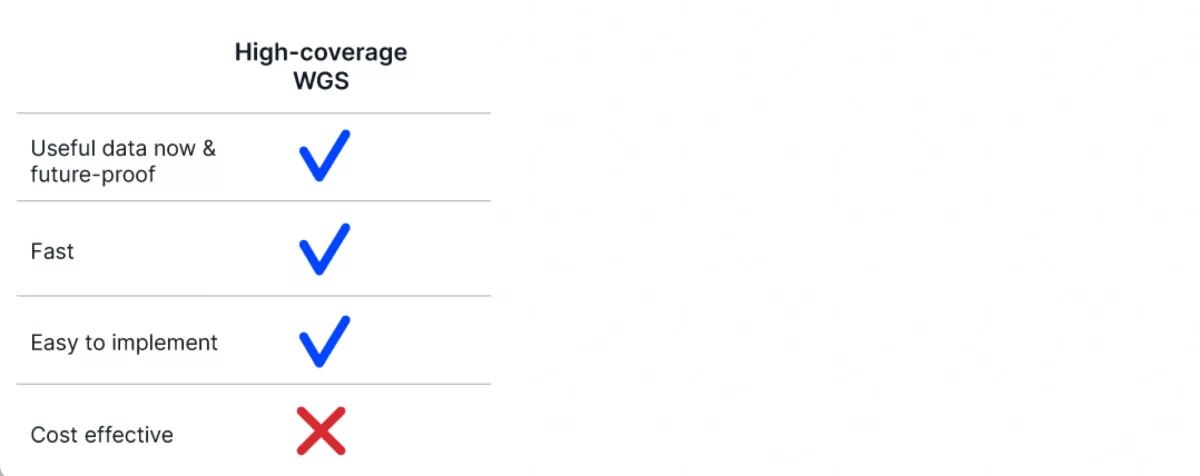
As sequencing technology has matured, it’s become a fast and easy data type to get your hands on. The instrumentation itself can be a little costly, but it is pretty straightforward to access a service provider or set up a sequencing machine in your own facility to capture the 30X coverage that is the gold standard of whole genome sequencing for variant detection projects.
Unfortunately, even though costs have been steadily declining over the past decade or more, sequencing to 30X coverage for every individual in a population is still prohibitively expensive. What this means is that high-coverage sequencing, while extremely useful, is still falling short of the requirements to be used as a routine breeding tool.
Microarray: the de facto tool for genotyping
For species that are well characterized with significant resources supporting the development of genomic tools, microarray has been a source of incredible breeding progress. Microarrays continue to be a go-to tool for genotyping populations for specific markers that aren’t expected to change much over time.
However, as thousands of samples get characterized over time, you tend to catch the outliers. Those genetic outliers often harbor useful genetic information that you want to bring into your breeding program and therefore screening tools. Unfortunately, adding new or additional markers to a microarray requires beginning the physical design process from the beginning, which is a significant time and financial investment.
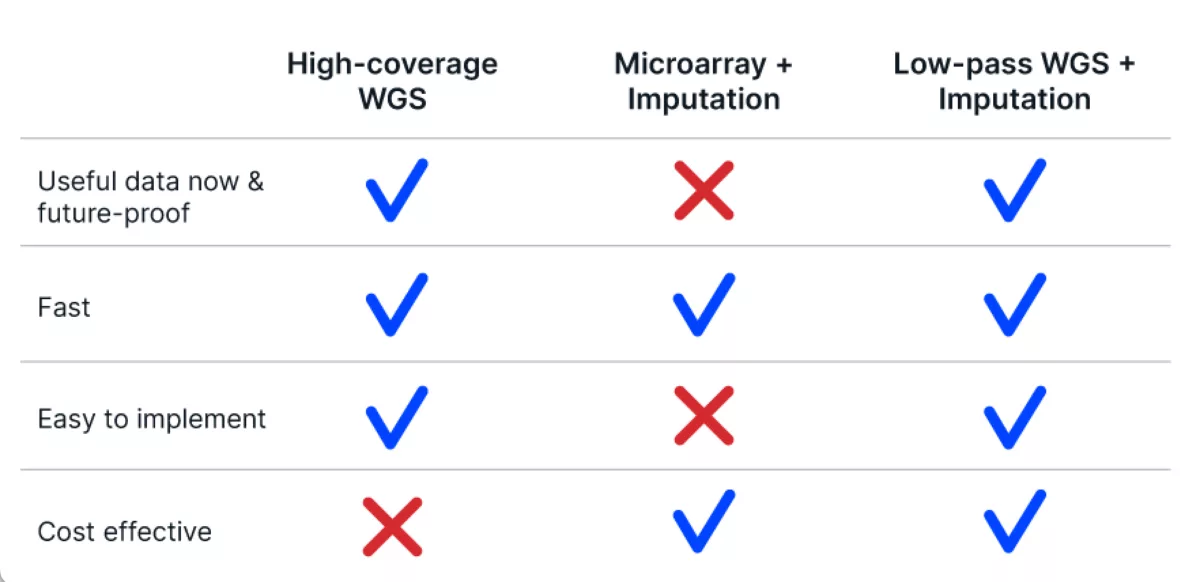
In addition, as time goes on, the important traits targeted by innovative breeding programs have increasingly complex genotype-phenotype relationships. Microarrays built on years or decades-old genetic information with a constrained number of set markers may not provide the power for genomic prediction that would move your program forward. So, while microarrays are fast and cost-effective, they lose their value over time and require significant resources to update.
Gencove: the modern approach to molecular breeding
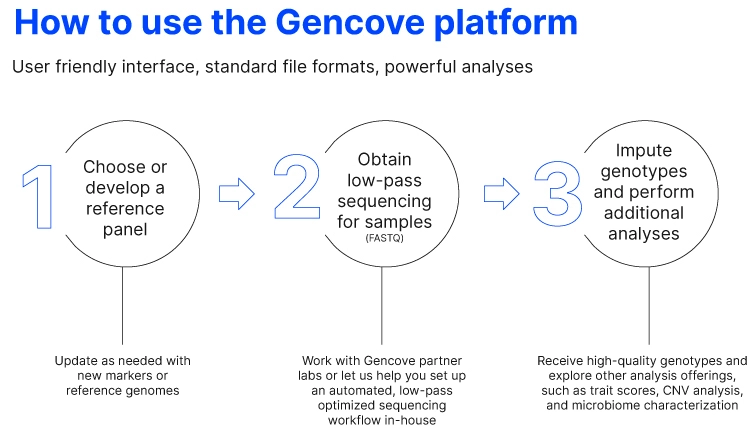
Thankfully, there is a method that meets all of the requirements. The Gencove platform leverages low-pass whole genome sequencing (typically <1X coverage), and purpose-built imputation software to provide accurate genotype information across millions of variants with genome-wide context.
Because the target coverage is typically less than 1X and takes advantage of an optimized library prep workflow, the method is cost-effective for even large sample volumes. Along with the specified markers that you’ve developed for key traits you need to breed, for now, the low-pass sequencing typically returns millions to tens of millions of variants across the entire genome that may be explored for later use as a marker.
Another advantage of a software-based genomics platform is the flexibility to update and backward compatibility that make it future-proof. As you explore new markers or develop new reference genomes, they can be incorporated into the genotyping pipeline for future screening, or previously sequenced samples can be re-run through the new pipeline.
Lastly, Gencove software is fast. You don’t have to sacrifice the turnaround time you’ve come to rely on from your other genotyping tools to capture the benefits of low-pass sequencing plus imputation.
The future of molecular breeding is squarely in sequencing technologies. As costs continue to decline, it becomes increasingly important to develop platforms that leverage sequencing data in useful, innovative, and cost-effective ways. Low-pass sequencing has a number of advantages over traditional genomics methods like genotyping arrays: tens of hundreds of times more data, and the ability to be implemented easily for a wide range of species and populations with limited set-up costs. The additional data results in increased statistical power to run genome-wide association studies (and identify genetic variants associated with a trait of interest) and/or rapidly deploy the technology in a manner that is customized to the population.