Biological sample and data management poses operational challenges at scale. Sample mixups and instances of file misnaming are almost inevitable when dealing with samples that have passed through a complex workflow, spanning multiple weeks and various parties.
We frequently encounter cases in which raw sequence reads coming off a sequencer have either been named improperly or inappropriately obfuscated compared to the original source's naming convention. Unfortunately, these discrepancies are often discovered only after analysis has already begun, leading to wasted compute time, manual intervention, and operational friction.
To address this, we are pleased to announce a new “sample manifests” feature within the Gencove platform, that catches and alerts to sample naming errors and will substantially decrease the incidence of these mix-ups. By default, as before, any FASTQs submitted to an existing Gencove project will immediately kick off analysis. However, we now offer the option to include sample manifest files in a Gencove project. These manifest files restrict the project to only accept FASTQs with "client IDs" that match those in a previously uploaded manifest file. FASTQs submitted with client IDs not found in the list will not be processed and will generate an error.
The sample manifest feature will aid in streamlining the process from biological sample to the solutions delivered by the Gencove platform by serving as a single “source of truth”, providing peace of mind that any results that are delivered are appropriately identified.
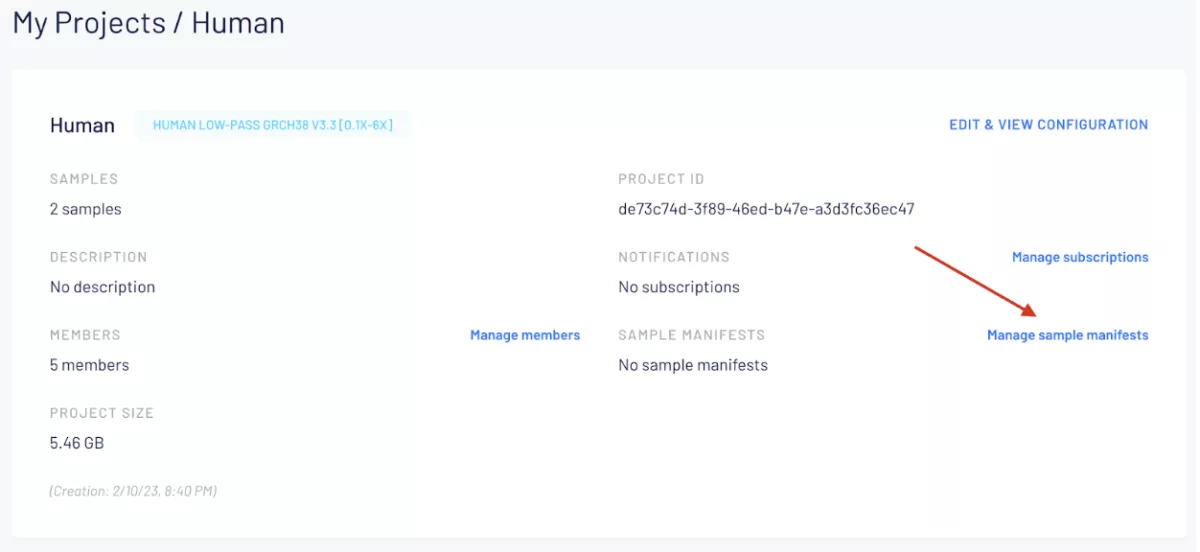
This feature is now live and can be enabled in any new or existing project by clicking the “Manage sample manifests” link in a project view. For more information, please see the full documentation at this link.