Users can now assign arbitrary JSON-formatted metadata to Gencove samples for downstream use.
We’ve been receiving consistent feedback from users saying they would like to store additional non-genomic data together with genomic data for their Gencove samples. These requests can be grouped into two broad clusters according to the nature of the data being stored:
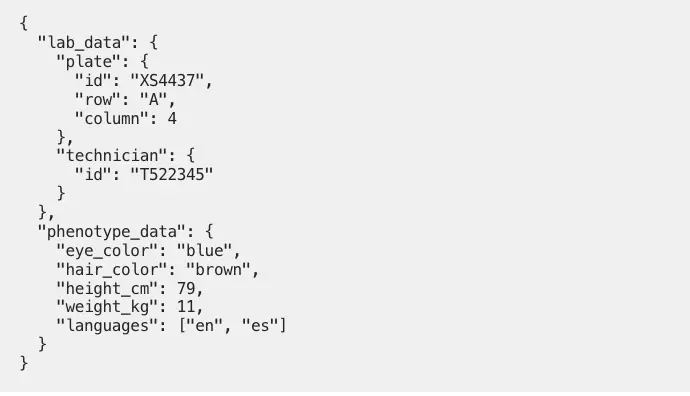
- Phenotypic data about the individual represented by the sample
- Technical processing data
- Batch information
- Alternative or auxiliary identifiers
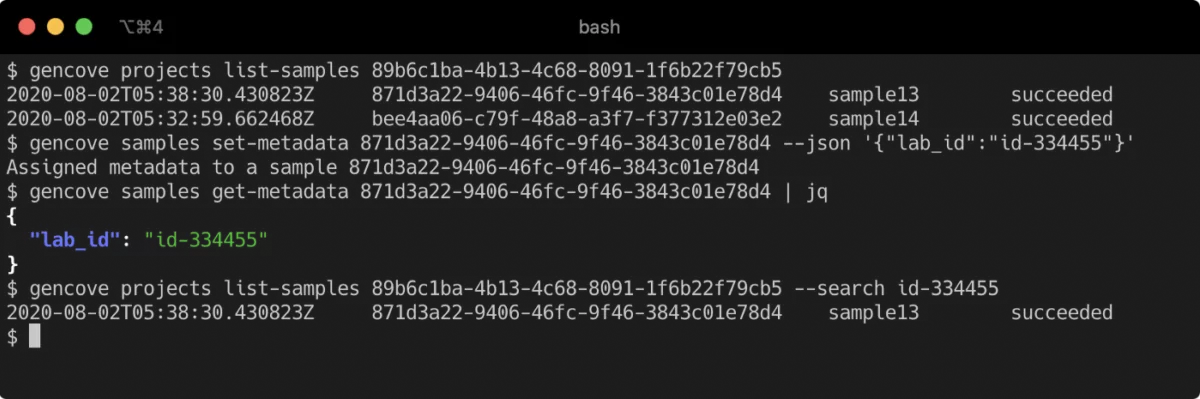
A commonality to these requests was also being able to search samples using sample metadata.